CRISPR Genome Engineering - gRNA Design and Analysis
Today we are announcing Benchling's platform for genome engineering using CRISPR.
Recent developments around CRISPR/Cas9 have taken the scientific community by storm and have important industry applications ranging from agriculture to gene therapy. We build powerful but easy-to-use software and want to do our part to propel this extremely promising field forward.
This post is the first in a series on genome engineering with Benchling. The first tool we're releasing in our editor is for finding gRNA target sequences in a region of interest and checking off-target binding.
Before we walk through this tool, note that it's now even easier to open a genomic region in Benchling. As we mentioned in our last post, Benchling's editor now supports sequences up to 5 Mb, and you can import in a chromosomal region from our import page. In our importer, the "Import from Database" option now supports gene names as well rather than just accession codes.
Finding gRNA target sequences
Inside our editor, gRNA design tools are located inside the genome engineering panel, activated by clicking the cross-hair icon in the right toolbar. After creating a new CRISPR design from here, you'll be able to set a region of the open sequence in which you'd like to find guides, as well as various parameters such as the PAM and guide length. Clicking "Search for Constructs" will show you all possible target sequences. The gRNAs will automatically show up on the sequence map to help you pick the ones in the right region. Benchling also supports paired design for nickase analysis.
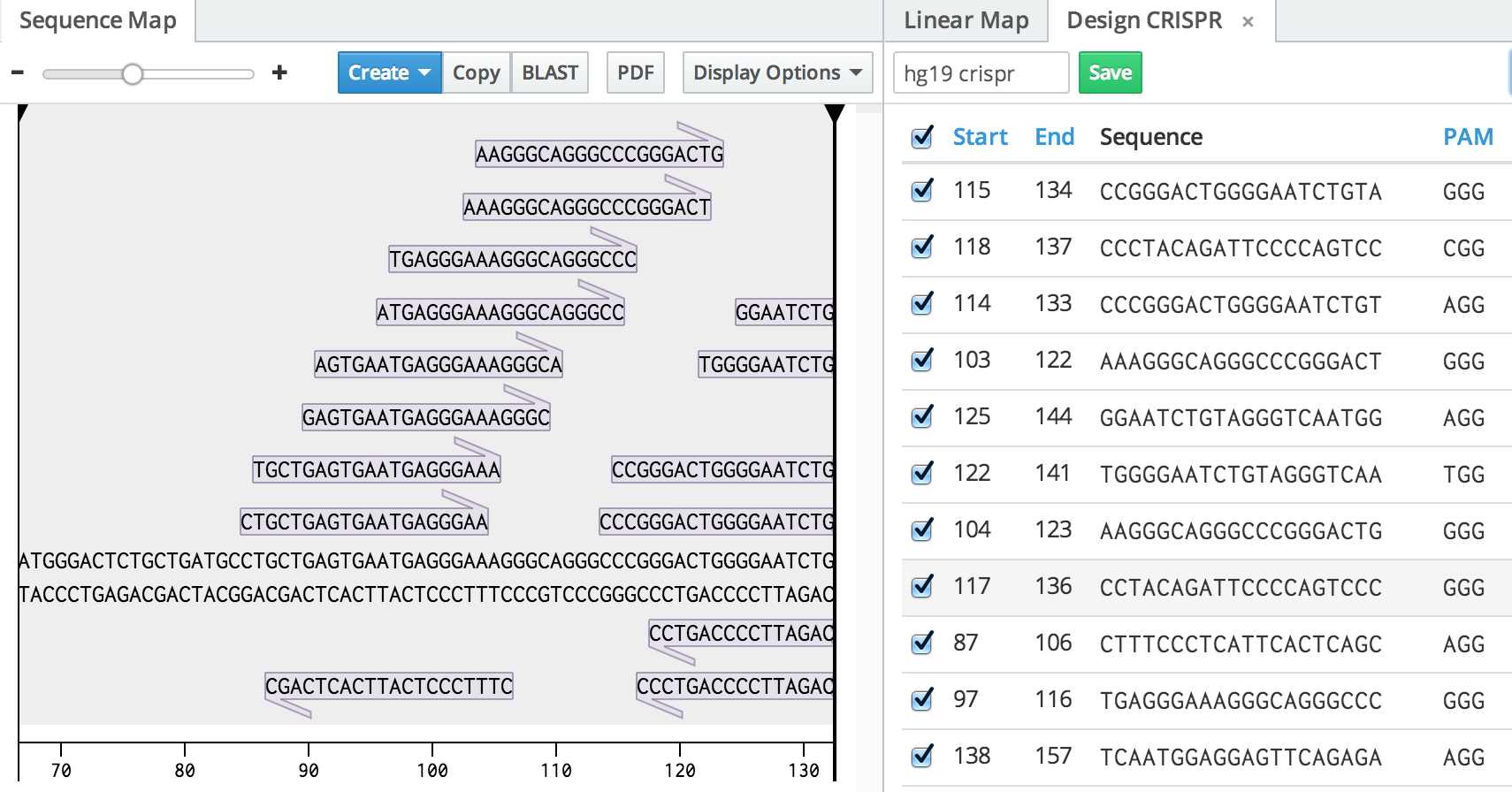
As with all of the analysis you do on Benchling, you can set a name and save your results in the top left of the CRISPR tab. It can then be re-opened from the genome engineering panel.
Checking for off-target binding
Benchling also provides fast and accurate off-target analysis for gRNAs. In the tab explained above, you'll notice that a genome can be set. There's also a "Genome Region" setting, through which you can input or find a region of the target genome that corresponds to your selected sequence, which removes it from off-target analysis.
Running the off-target analysis for ~20 guides should reliably complete in just around a minute. Behind the scenes, we index gRNA sites for supported PAMs and genomes to make these searches efficient without losing accuracy.
The guides are scored by their off-target effects using a method developed by the Zhang lab at MIT. Clicking on each score pulls up the location of each off-target site, along with links to the UCSC genome browser and a gene if the site occurs in an exon.
Saving gRNAs
After you've picked gRNAs that target the correct location and don't exhibit problematic off-target effects, the CRISPR design tab allows you to save those to a primer library, and automatically attaches them to the sequence you are viewing. You can easily share the primer library with colleagues and export them to CSV for ordering.
What's Next
We have extensive plans for our genome engineering platform. In the short term, look for features like automatic design for PCR primers to verify gRNA activity and integration with the Assembly Wizard to automate in silico insertion of gRNAs into reference CRISPR plasmids.
We have an efficient pipeline for processing and storing genomes, so if there are new genomes you would like to see added or you are interested in working with custom genomes, drop us a line at contact@benchling.com.
Powering breakthroughs for over 1,200 biotechnology companies, from startups to Fortune 500s
