Edit single bases with Benchling!
Currently, scientists can edit genes, with CRISPR, but how well can this method edit a specific base? Not very well — only 0.1% to 5%, and at best, 20% [1-3].
That’s all about to change! Alexis Komor and others from David Liu’s lab at Harvard recently devised a CRISPR mechanism, called base editor, that specifically edits single bases with correction efficiencies of 15%-75%, alongside indel formation of only 0.1%-5% [4].
If you want to design your own gRNAs to base edit, you can do it on Benchling. The tool will identify possible gRNAs for your target region, and provide you information about the gRNAs’ efficiencies and off-target scores. A big shoutout to Alexis Komor and Ben Thuronyi from the Liu Lab, who helped co-develop this feature.
Step 1. Identify the genomic region you want to target
Start by clicking “Create,” and then click “Create CRISPR guides”. In the next dialog box, you can search for your gene of interest. Here is what it looks like to search for BRCA2 in a human genome (hg38).
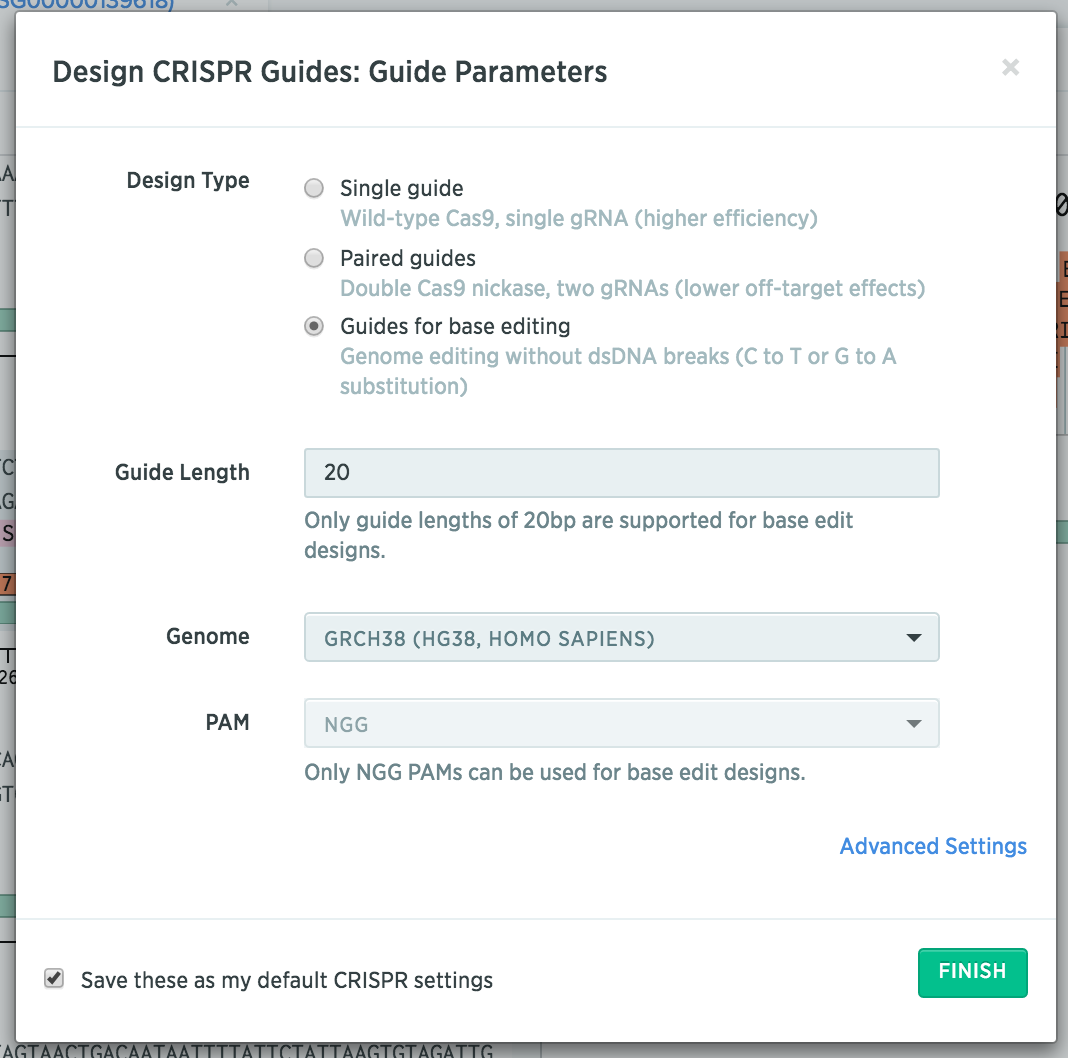
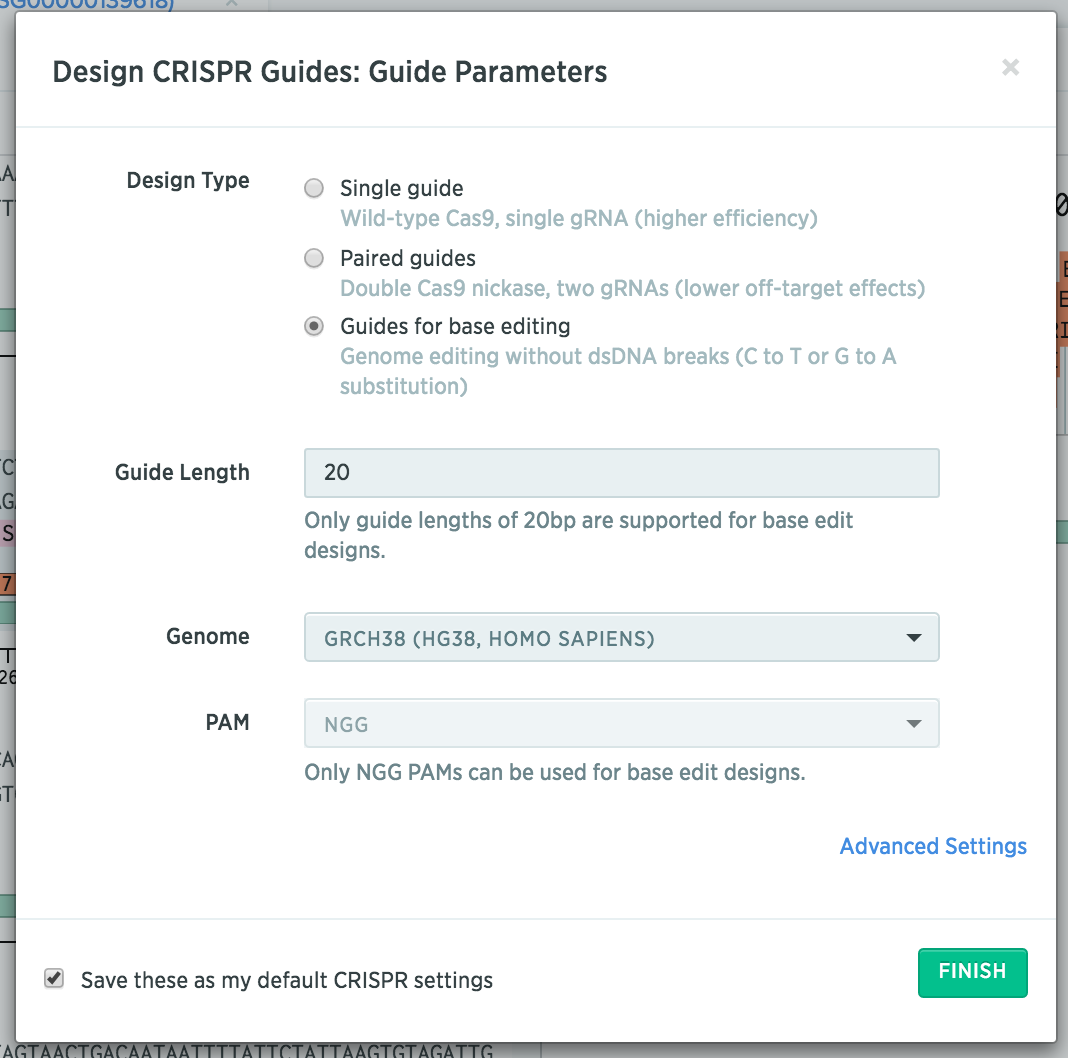
After importing your gene of interest, select “Guides for base editing.”
Tips
We support over 50 different genomes! If your favorite genome is not on the list, let us know, and we can import it for you.
The base-editing score is calculated based on the BE1 enzyme, so if you decide to use BE2 or BE3, the efficiency could be much higher than the score indicated [4].
Step 2. Find all possible gRNAs for your target site
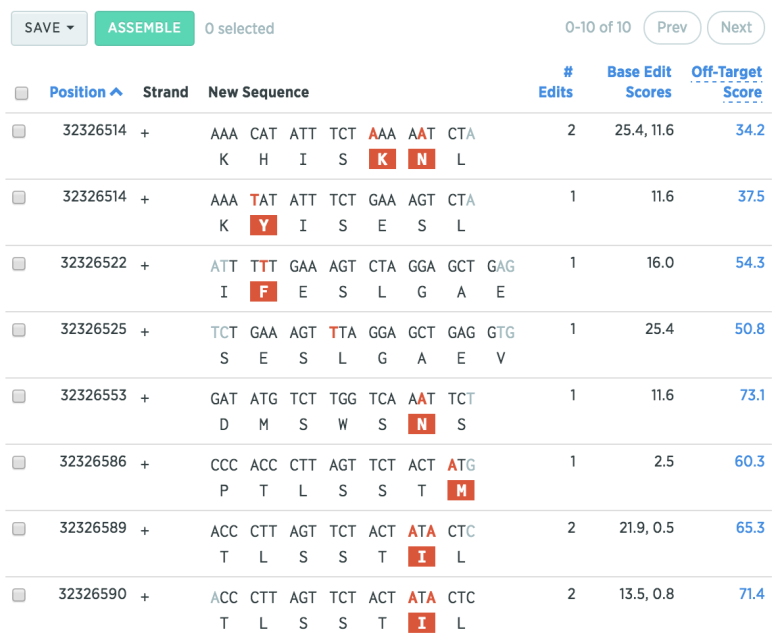
Select a target region (such as Exon 7), and hit the “+” button to find possible gRNAs for that region. Benchling will show where the point mutation(s) will occur, along with the base-edit score (efficiency), and the off-target score.
Tip
You can sort your list of gRNAs by clicking on the section headers (e.g. “# Edits” or “Off-Target Score”)
Step 3. Select a gRNA with high base editing efficiency and low off-target rates
You’ll want to choose a gRNA with a high base editing score, and a high off-target score. The base editing score refers to the mutation efficiency, which is calculated based on the sequence surrounding the target region [4]. The off-target score is calculated according to the inverse probability of Cas9 off-target binding [5]. A higher score means the sequence has a smaller chance of binding to sequences in the rest of the genome.
Tips
If translations are not automatically imported with your gene, you can view the translational mutations by selecting the targeted region’s open reading frame, right-clicking “Create Translation” and then “Forward”.
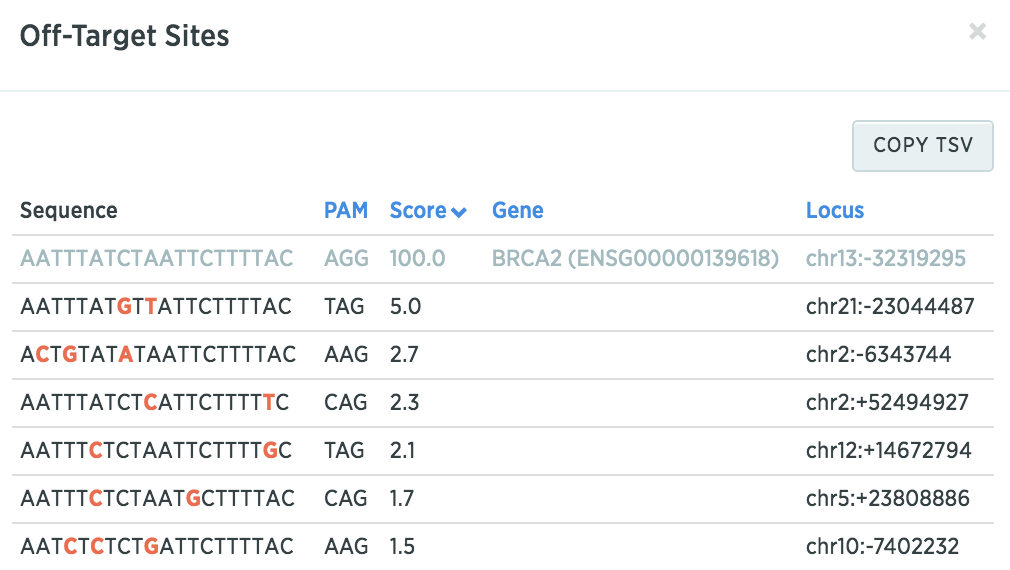
You can click on the off-target score to see potential off-target sequences to determine if these sequences are part of the genes relevant in your studies (mismatches are denoted in red).
What’s next?
There are a few ways you can go about expressing your chosen gRNA:
Use Dr. Komor’s vector and then perform a PCR/ligation procedure
Perform in vitro transcription
Choose from a preloaded expression vector (or upload your own vector) by clicking “Assemble” from the gRNA list
As for deciding between BE1, BE2, and BE3, Dr. Komor recommends using BE3 for most applications. If low amounts of indel formation are not acceptable, then she recommends using BE2. The plasmids can also be obtained from Addgene.
Other ways to design CRISPR experiments on Benchling
We’ve developed a whole set of tools on Benchling that you can use for your CRISPR experiments! Take a look at some of our previous blog posts to see how Benchling can best suit your CRISPR needs.
References
Maruyama, T. et al. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nature biotechnology 33, 538-542, doi:10.1038/nbt.3190 (2015).
Wang, H. et al. One-Step Generation of Mice Carrying Mutations in Multiple Genes by CRISPR/Cas-Mediated Genome Engineering. Cell 153, 910-918, doi:10.1016/j.cell.2013.04.025 (2013).
Mali, P. et al. RNA-Guided Human Genome Engineering via Cas9. Science 339, 823-826 (2013).
Komor, A. et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420-424 (2016).
Hsu, Patrick D et al. "DNA targeting specificity of RNA-guided Cas9 nucleases." Nature Biotechnology 31.9, 827-832 (2013).
Powering breakthroughs for over 1,200 biotechnology companies, from startups to Fortune 500s