The Impact of mRNA Diversity on Protein Expression
Contributed by:

STEPHEN FLOOR, DOUDNA LAB, Helen Hay Whitney Postdoctoral Fellow in the Doudna Lab
Editor's note: At Benchling, we are dedicated to helping create and communicate cutting-edge research. We launched this site in conjunction with Dr. Stephen Floor and Dr. Jennifer Doudna's publication in eLifein 2016. If you want a similar website for your research, just contact us.
Each human gene on average can be transcribed into five different kinds of messenger RNA products (so-called transcript isoforms). This raises an important question for cell biology: with all those diverse mRNAs present, how does a cell control protein translation? Scientists Dr. Stephen Floor and Dr. Jennifer Doudna developed a technique (TrIP-seq) to study the relationship between transcript isoforms and translation. Eventually, they understood translational control well enough to graft different components in transcripts to tune the protein expression of a transfected foreign gene.
In this article, Dr. Floor joins Benchling to help us understand how he and Dr. Doudna tackled the problem of mRNA translation. He also provides a step-by-step TrIP-seq protocol in Benchling and information on how to search translation level of all transcript isoforms from their results. Their research helped us understand translational control in human cells better and now scientists can use a similar approach to understand translational control in other cell types.
What is a transcript isoform?
Eukaryotic genes often express more than one type of messenger RNAs. These mRNA types, called transcript isoforms, are built by alternative transcription, splicing, or polyadenylation of a single gene. In humans, an average gene expresses five different isoforms, but some have as many as 80 (see below, Ensembl GRCh37 annotations). Transcript isoforms can contain different coding or regulatory sequences, or both. Each of these transcript isoforms can therefore perform different functions. For example, you can determine whether a protein will be shuttled to the membrane or retained in the cell by changing the 3'UTR of the transcripts (Berkovits & Mayr, 2015). Changes in transcript isoforms are also associated with many human disorders, such as muscular dystrophy, amyotrophic lateral sclerosis (ALS), and diverse cancers (Scotti & Swanson, 2016).
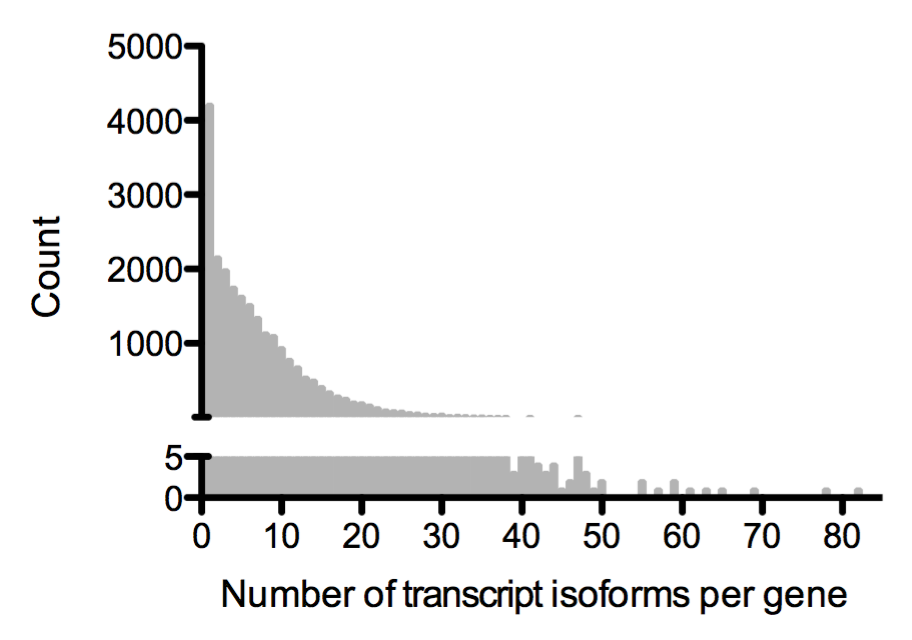
How do different transcript isoforms influence gene expression?
It is hard to predict protein levels based on RNA expression. Messenger RNAs contain different sequences that define how well the mRNA is translated, and therefore how much protein is made from the open reading frame (ORF). Different regulatory sequences, such as 5' leader sequences, upstream ORFs, microRNA binding sites, AU-rich elements, and sequences around start codons (Kozak context) can all change the protein production level of the ORF in eukaryotes. As different transcript isoforms can have a different complement of regulatory sequences, they may be differently translated. Therefore, to understand the relationship between RNA and protein expression, we need to know the translation levels of different transcript isoforms. Different ORFs be uncovered using ribosome profiling (Ingolia, 2014), but untranslated regulatory sequences are invisible to this technique.
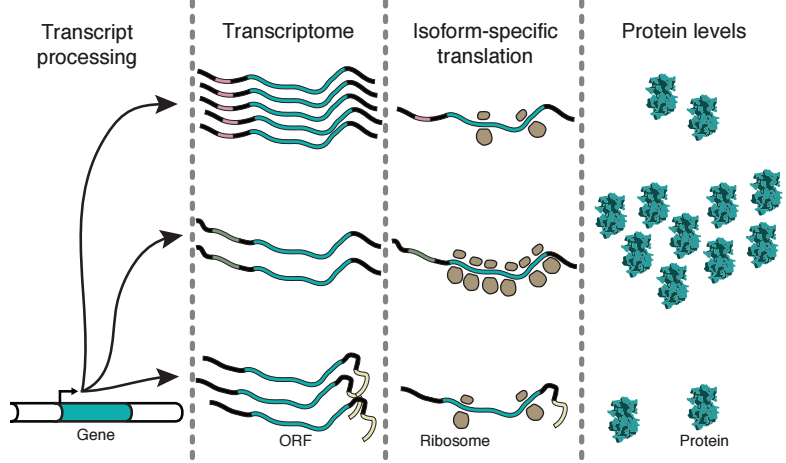
Floor & Doudna's solution: Measure the gene expression of different transcript isoforms (TrIP-seq)
Multiple ribosomes can associate with a single mRNA, which is known as a polysome. Polysomes can be separated into peaks containing different numbers of ribosomes using high-speed centrifugation. We reasoned that we could estimate the translation level of each transcript isoform by determining how many ribosomes each isoform is bound to using polysome fractionation and RNA-seq. We termed this approach Transcript Isoforms in Polysomes sequencing (TrIP-seq), which is described in full in our paper.
Here is a flowchart of the experimental procedure of TrIP-seq:

We provide a comprehensive protocol of this method of extracting mRNAs bound to different numbers of ribosomes here. You can use this method to measure the translation levels of transcript isoforms of your interest. The protocol can be saved as PDF, or you can edit your copy directly within Benchling.
Download TrIP-seq protocol here.
Floor & Doudna's database: Translation level of all expressed transcript isoforms
We have used TrIP-seq to measure the translation levels of all expressed transcript isoforms in human HEK 293T cells. We also demonstrated that we could tune the protein expression level of an arbitrary gene based on what we learned from our data. If you want to see how transcript isoforms impact the translation of your favorite gene in human cells, you can search for your gene here.
For a full description of the technique and conclusions, features of mRNAs that cause differential translation, and how to use these data to predictably control translation of orthogonal genes, please see Floor & Doudna, eLife, 2016.